Introduction
 In my previous post I explained the basics of genetic algorithms, and provided a very simple example written in Objective-C. In this post I will dive deeper into the fascinating world of genetic algorithms and show a more sophisticated solution to a more difficult problem, known as the 8 Queens puzzle.
In my previous post I explained the basics of genetic algorithms, and provided a very simple example written in Objective-C. In this post I will dive deeper into the fascinating world of genetic algorithms and show a more sophisticated solution to a more difficult problem, known as the 8 Queens puzzle.
The demo program can be downloaded from github here.
Debt of gratitude
My study of genetic algorithms has been largely based on reading blog posts written by other developers. The post I found most helpful, and which inspired me to tackle the 8 Queens puzzle, can be found on the HandCraftsman blog (a.k.a. Clinton’s Blog). He has posted two separate solutions to this puzzle, one written in C# and the other in some language called Go. The post I studied the most, and from which my algorithm’s gene encoding system is based, can be found here. My algorithm is heavily influenced by Clinton’s but different in several ways.
The 8 Queens Puzzle
Back in 1848 some chess guru named Max Bezzel posed the question of how can you arrange eight queens on a chessboard such that none of them can attack each other. Some seriously smart people, including the almighty Gauss himself, gave this problem a lot of thought and it soon became a well-known problem amongst mathematicians. These days it has become a common problem for Computer Science students to solve while learning about things like constraint programming, logic programming, or genetic algorithms.

Learn more about the 8 Queens puzzle here.
Not the fastest draw in the West
I should point out that genetic algorithms are not the fastest way to solve the 8 Queens puzzle. There are many other ways to solve it, some of which can find all ninety-two distinct solutions to the puzzle in less than one second. It took my algorithm just under an hour on a decent MacBook Pro to find all ninety-two distinct solutions.
I’m OK with that. I didn’t set out to write the fastest way to solve the 8 Queens puzzle. Instead, I used the 8 Queens puzzle as an interesting problem with which I could explore designing and implementing genetic algorithms.
From Hello World to 8 Queens
In my introductory post about genetic algorithms I reviewed a very simple example; an algorithm that evolves randomly generated strings to match a string typed in by the user. In that algorithm the genetic sequence stored in a chromosome has a direct, literal relationship with the problem domain. In other words, the characters/genes in the string/chromosome can be compared to the user input string to see if the chromosome is an exact match. In that contrived example the genes are not being used to encode information about a problem domain: they are that information.
In such a simple example it was not possible to create a genotype-phenotype distinction. A genetic algorithm that solves the 8 Queens puzzle, on the other hand, is a step up in complexity. Genes representing a possible solution to the 8 Queens puzzle will be meaningless by themselves. They must be projected, or, to stick with terms used in biology, translated, into information about the problem domain. This is similar to how nucleotides in DNA are translated into functional proteins within your cells.
Learning from my mistakes
Designing an appropriate genetic encoding system, and determining what information the genes should encode, is of crucial importance to creating a successful genetic algorithm. I know this from experience. My first attempt didn’t work out. I am going to explain my faulty reasoning, and what I learned from it, to better illustrate why the encoding system I eventually created works well.
My initial thought about how to encode information to solve the 8 Queens puzzle was to have a gene sequence consisting of sixty-four genes, since there are 8×8 squares on a chessboard. Each gene would represent one square of the chessboard and encode the binary state of a square (queen/no-queen). Each gene would be stored as a character in a string. The first gene would represent the top-left square on the chessboard, the ninth gene would represent the first square in the second row, the last gene would represent the bottom-right square, etc.
I started implementing a genetic algorithm using this encoding system, but soon ran into a problem indicative of a serious design flaw. My code needed to keep enforcing the rule that each chromosome must have exactly eight genes that represent a queen. This problem appeared when creating a new chromosome with randomly generated genes, and when combining the gene sequences of two mating chromosomes.
The problem around mating chromosomes and ensuring a child’s genes contained exactly eight “queen genes” was particularly troubling. Adding and removing queen genes willy-nilly from each new gene sequence meant introducing a lot more randomization into the algorithm. Genetic algorithms rely on randomness for both creating the initial population’s gene sequences and introducing mutations to subsequent generations, but those are highly controlled and intentional randomizations.
The algorithm’s crossover method, which is the fancy name for how it combines segments from two gene sequences to form a new one, should not introduce a high degree of randomness that significantly alters the combined gene sequence. Doing so renders the algorithm’s fitness function pointless. What’s the point in mating the fittest chromosomes if you always make a large number of random modifications to their children?
The fundamental problem with my initial encoding system was that it stored irrelevant information about the problem domain. It encoded the state of all sixty-four squares on the chessboard. Producing new gene sequences involved a lot of clean up work, which introduced inappropriate randomization. A well-designed encoding system would not include superfluous information about the chessboard.
Designing a proper encoding system
A solution to the 8 Queens puzzle can be expressed as a set of eight data points. Each data point represents a square of a chessboard on which a queen exists. Therefore, each chromosome used in a genetic algorithm that solves the 8 Queens puzzle should consist of eight genes, each of which identifies a square of a chessboard in which a queen exists. Rather than store the binary state of all sixty-four squares, the gene sequence should store only the locations of the queens.
My genetic algorithm represents the top-left square on the chessboard with the character zero (‘0’) and the bottom-right square with lowercase p (‘p’). This is a range of sixty-four characters, according to the ASCII character codes. In other words, ‘p’ – ‘0’ is equal to 64. My choice of using the character for zero as the initial character is mostly arbitrary, but I liked the idea of the first square being represented with a zero (as in, squares[0]).

In the demo program, there is a class that is used to store a solution to the 8 Queens puzzle and draw a diagram of it to the console. An example of its output is seen below:
. Q . . . . . . . . . . . Q . . . . . . . . . Q . . Q . . . . . Q . . . . . . . . . . Q . . . . . . . . . . Q . . . . . Q . . .
That class is named JAS8QueensSolution. It contains the following methods that draw the solution to the console window. This code shows how information from the problem domain (squares on a chessboard) can be translated via the genetic encoding system to check for the presence of a queen at a particular chessboard square.
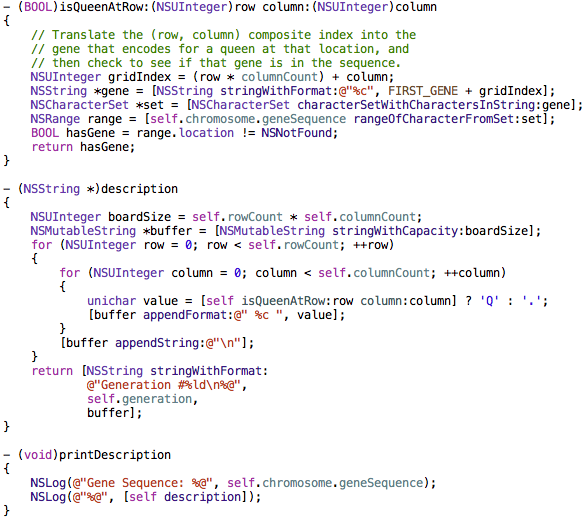
The genetic encoding system is also used when evaluating the fitness of a chromosome’s gene sequence, which we will look at next.
Working out the fitness function
In my previous blog post about genetic algorithms the demo program’s fitness function was very simple. It took a chromosome (which simply wraps a string) and compared each character/gene against the corresponding character/gene from the target string. The greater the delta between the two characters/genes, the lower the chromosome’s fitness became. A chromosome with a fitness of zero meant it was an exact match to the target string; anything below zero meant the chromosome was not an exact match.
That simplicity is not possible when genes do not directly map to, or represent, the problem domain. A level of abstraction exists which allows a gene in the 8 Queens algorithm to encode information about a square of a chessboard on which a queen exists. As such, the fitness function for my 8 Queens genetic algorithm must translate each gene into information about the problem domain while evaluating a chromosome’s fitness.
An interesting quirk about the 8 Queens puzzle is that the other genes in a chromosome determine a given gene’s fitness. A gene represents the location of a queen, and a queen’s safety is contingent upon the locations of the other queens. It turns out that the fitness function can be significantly optimized due to this quirk. Instead of searching for queens occupying squares that are North, Northeast, East, Southeast, South, Southwest, West, and Northwest of a given queen, we only need to search half of those directions (the demo program searches North, Northeast, East, and Southeast.) For example, if a queen is South of another queen, searching North from both of those queens will cause the incompatible locations to be detected. There is no need to search South as well.
Here is the fitness property from JAS8QueensChromosome that calculates the chromosome’s fitness.

That property calculates the fitness of each gene and caches the sum. The result is cached to improve performance, since this property can be accessed multiple times for a chromosome. The helper method that calculates the fitness of a given gene is seen here:

This method searches in four directions, as discussed earlier, to look for other queens that are attacking the queen represented by the gene at the given index. Let’s take a look at the code that searches North and East for attacking queens. This code shows how the genetic encoding system is used to translate genes into information about the problem domain, namely, the squares in which queens exist.

The translation of genetic information to problem domain information occurs in the for() loops. The ‘threat’ variable in those loops identifies a chessboard square. A list of chessboard square identifiers is appended together and passed to the areEmptyThreats: method for analysis. That method checks to see if the chromosome’s gene sequence contains any of those chessboard square identifiers. If so, that means a queens exists on one or more of those squares.
Breeding the next generation
Because each chromosome contains exactly the information needed to represent a solution to the 8 Queens puzzle, the code to mate two chromosomes is very straightforward. All of the problems I faced with my original genetic encoding system, as described earlier, disappeared when I used the system seen above. Here is the code from JAS8QueensChromosome that mates one chromosome with another:

The chromosomes are mated after the algorithm has evaluated the fitness of every chromosome in the population to look for valid solutions to the 8 Queens puzzle. Below is the code from JAS8QueensAlgorithm that selects the best chromosomes from the population, along with a small number of the non-elite, and creates the next generation.

Wrapping up
There is a lot more going on in the demo program than what I’ve shown here, so I encourage you to grab it off github and take a look. One particular point of interest is how JAS8QueensChromosome overrides the hash and isEqual: methods to help the algorithm avoid adding equivalent chromosomes to the population.
The demo program can be downloaded from github here.
Thanks for stopping by!


Pingback: Simple Genetic Algorithm in Objective-C | iJoshSmith
Thanks a lot for this post, Josh!
you’re saving my ass with your genetic algorithms implementations! haha
hope we can keep in touch!
Thanks a lot.
Thanks, I’m glad you like it.